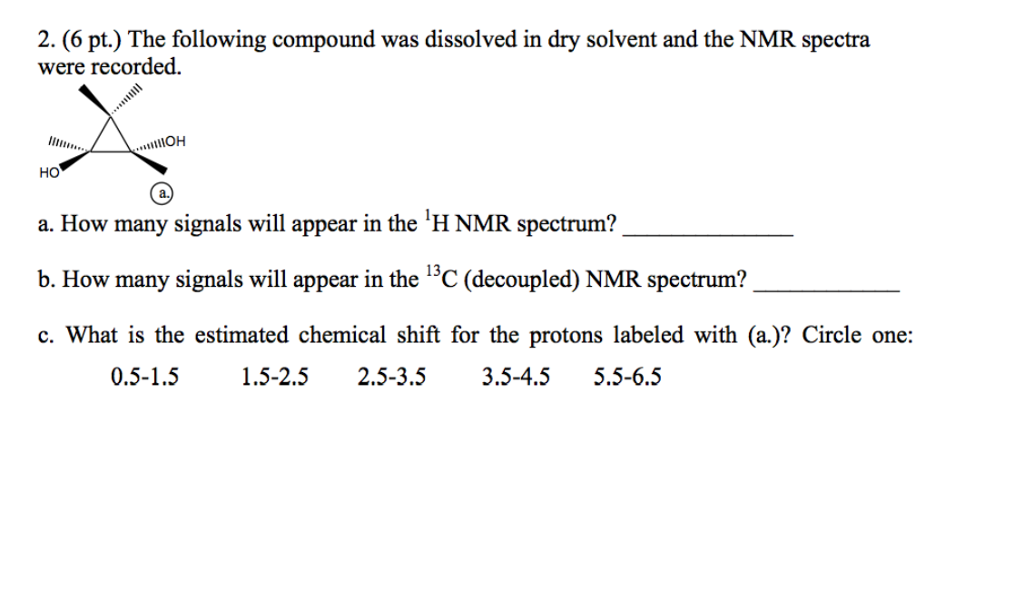
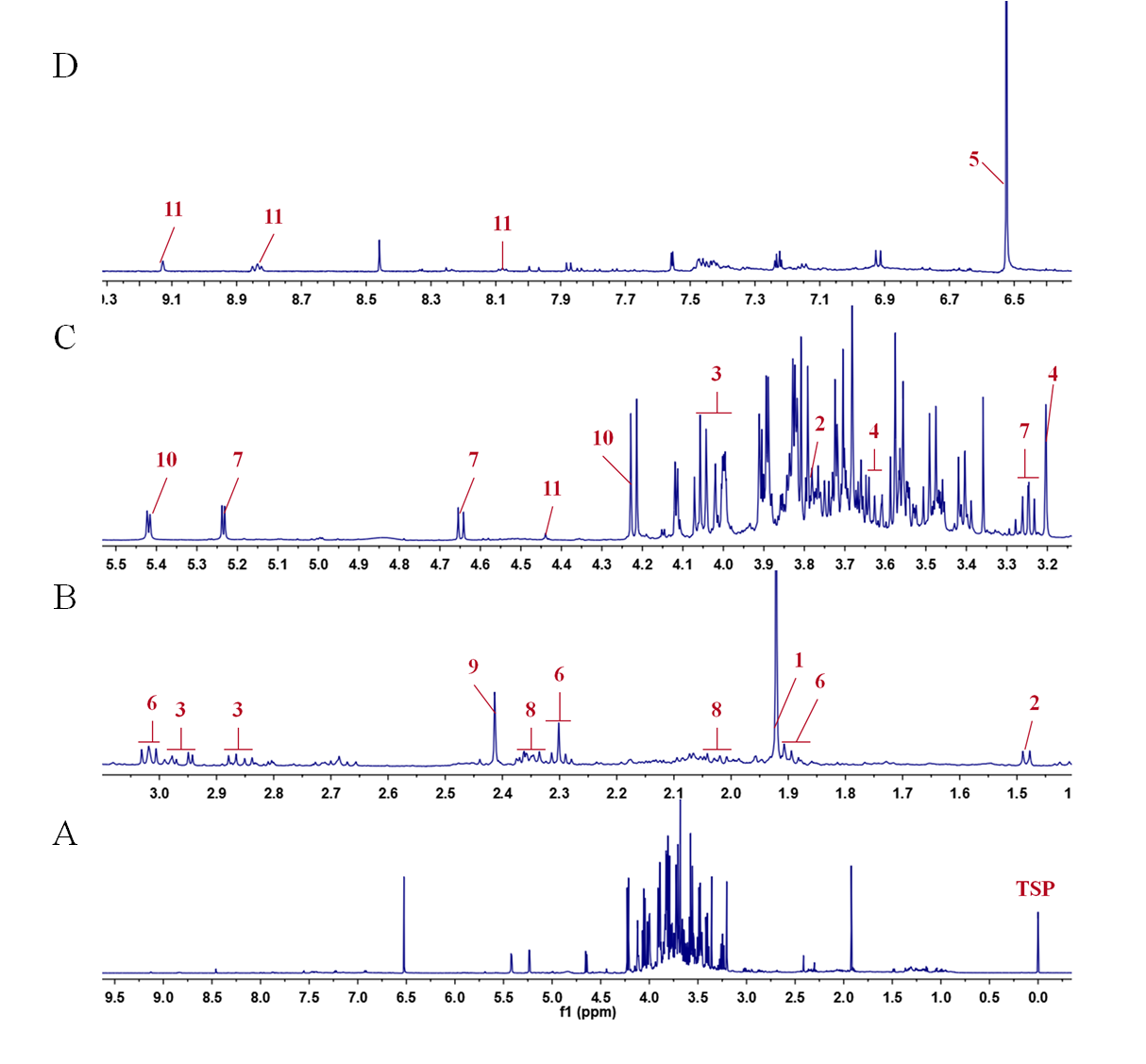
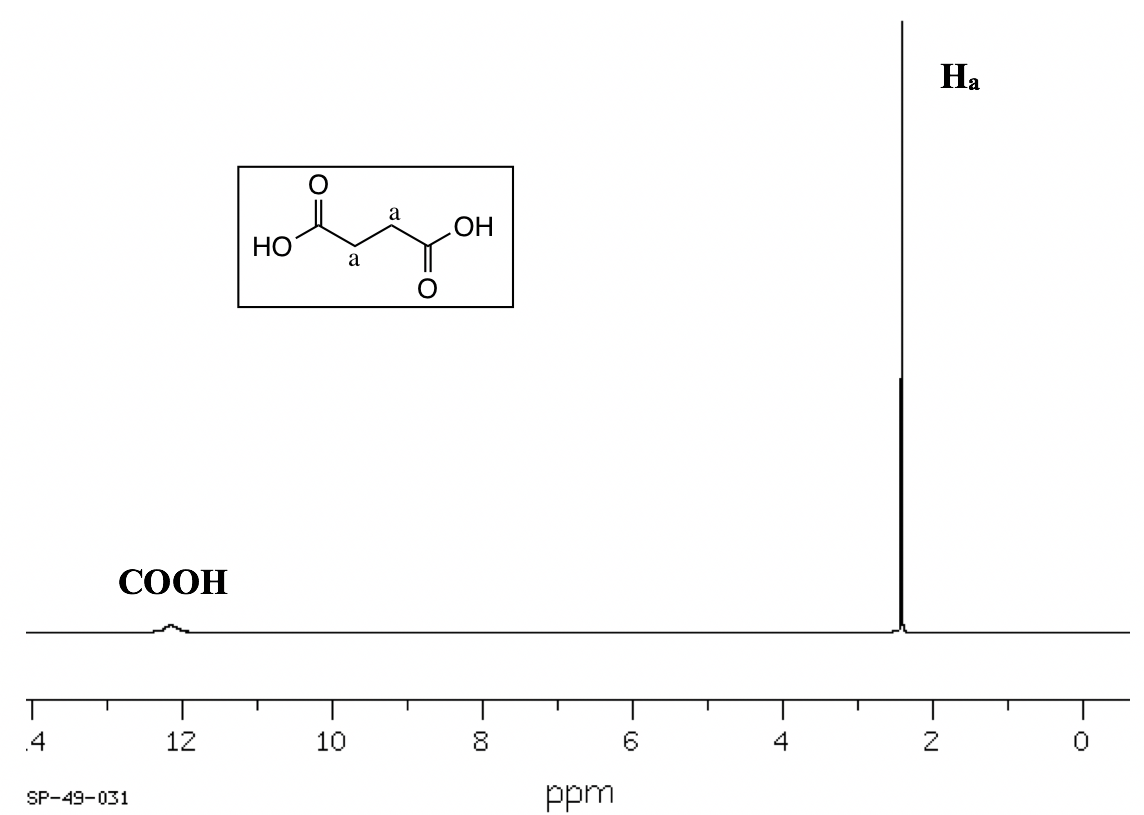
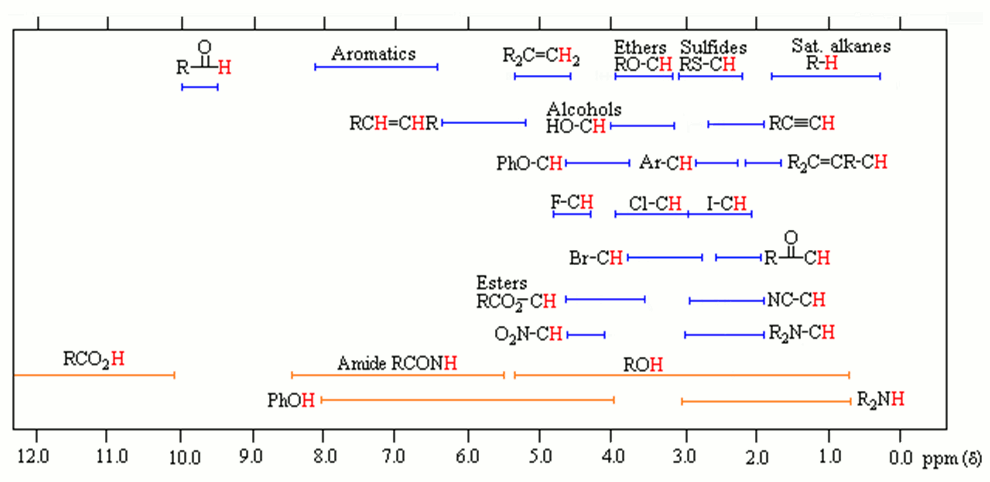
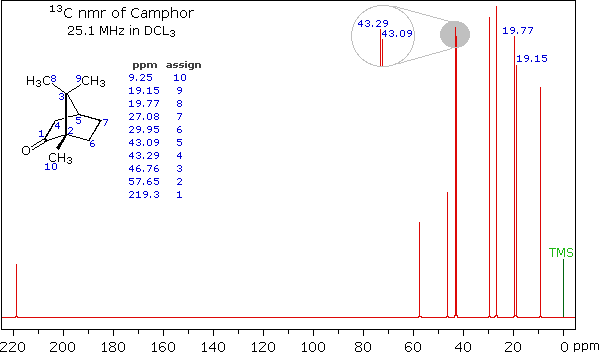
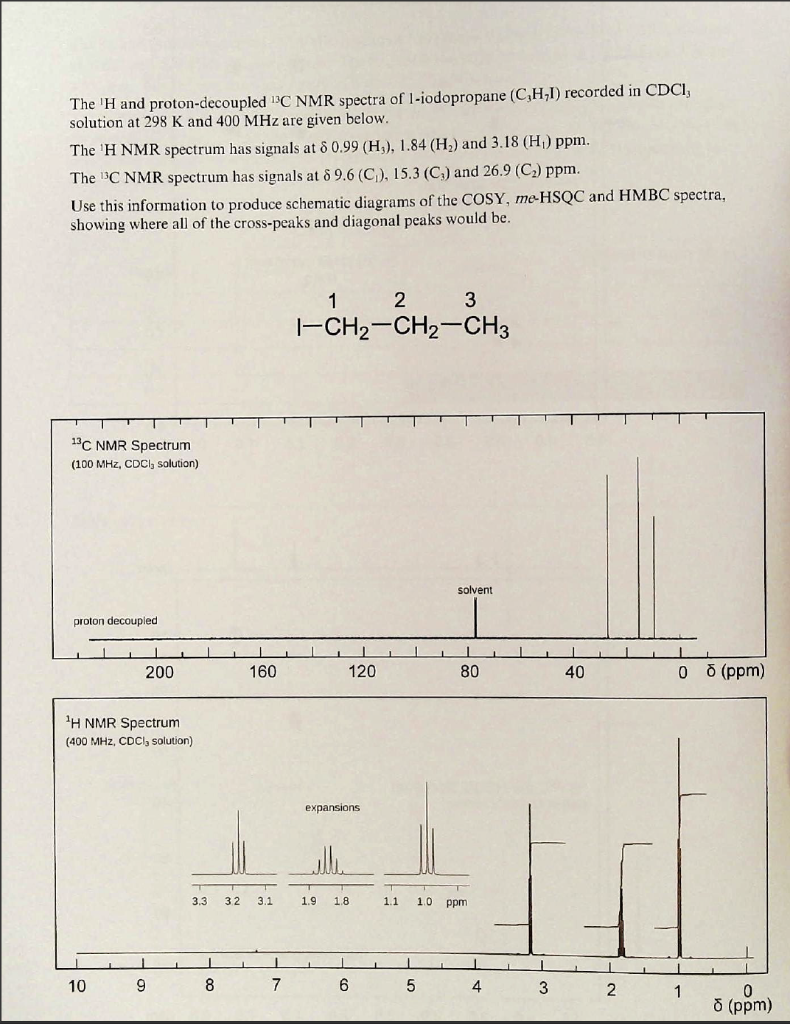
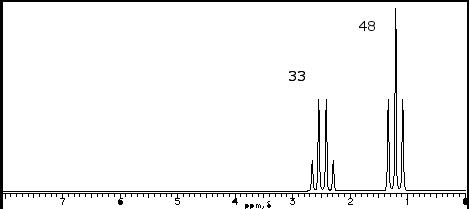
SOLVED: 1. The 1H NMR, C1H14O3NBr is provided_ data for an unknown compound R with the formula (b) The sample was dissolved in CDClz and all NMR spectra were recorded on Bruker
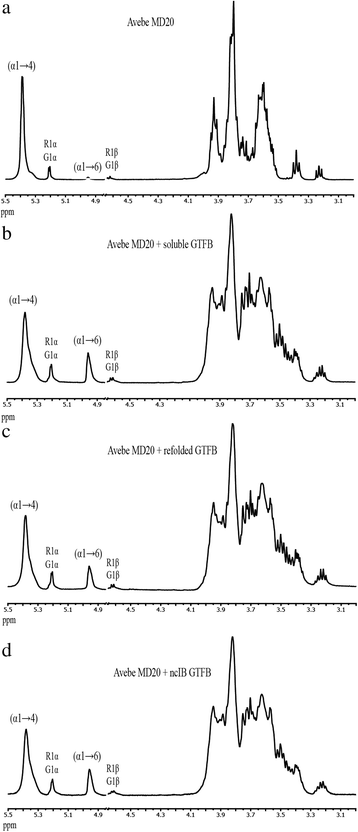
Characterization of the 4,6-α-glucanotransferase GTFB enzyme of Lactobacillus reuteri 121 isolated from inclusion bodies | BMC Biotechnology | Full Text

The 500 MHz 1H-NMR spectra were recorded using a protein concentration... | Download Scientific Diagram

1 H-NMR spectra were recorded at 300 MHz of a pH 9.0 solution D (0.020... | Download Scientific Diagram
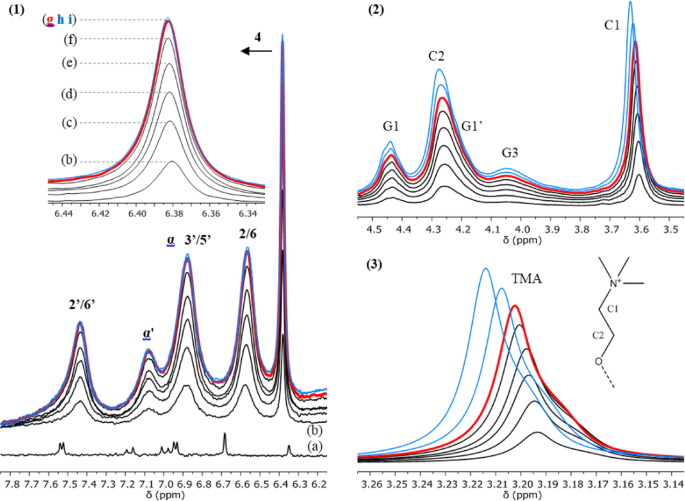
1H NMR study of the interaction of trans-resveratrol with soybean phosphatidylcholine liposomes | Scientific Reports
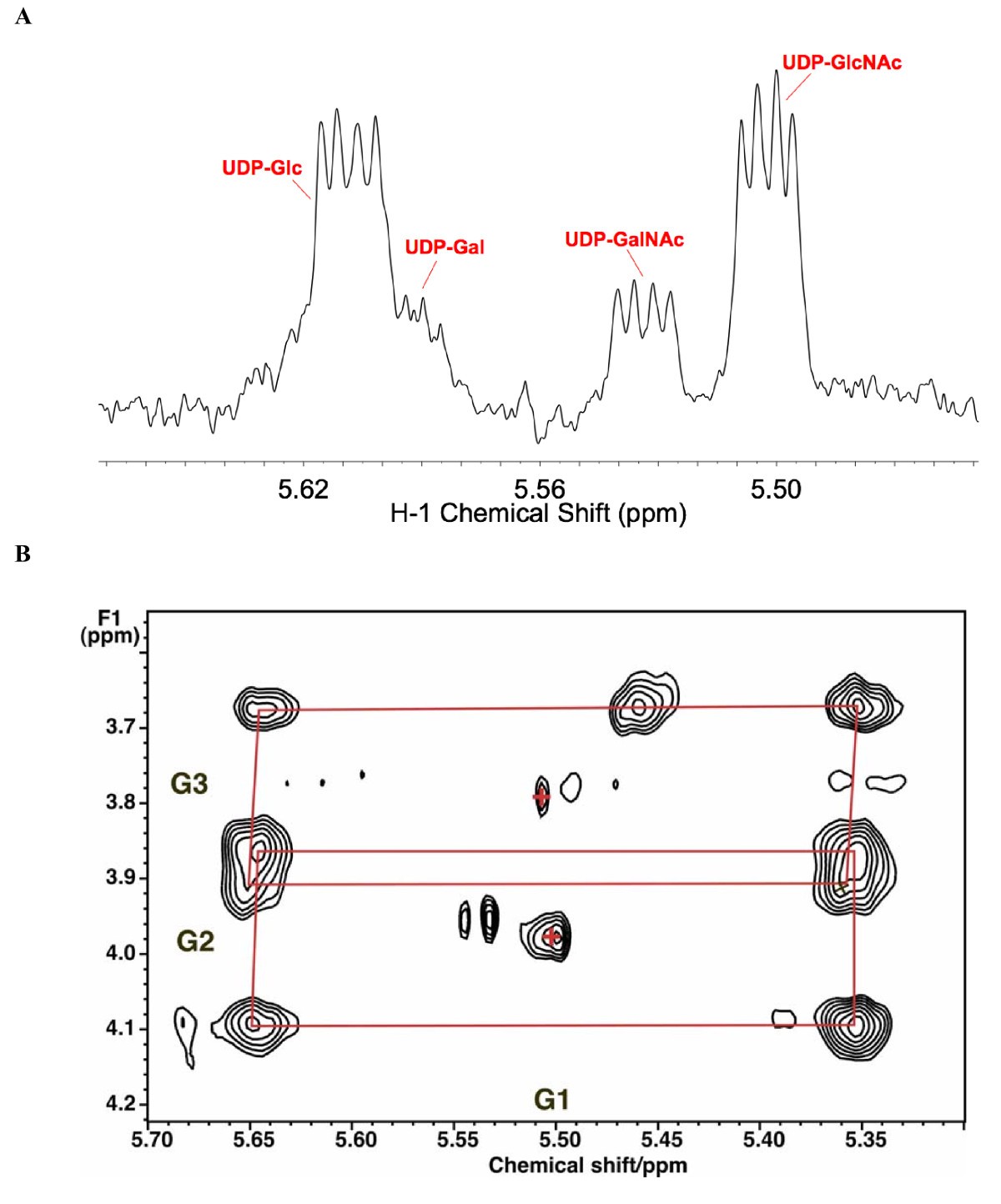
A novel deconvolution method for modeling UDP-N-acetyl-D-glucosamine biosynthetic pathways based on 13C mass isotopologue profiles under non-steady-state conditions | BMC Biology | Full Text

A Solution NMR Approach To Determine the Chemical Structures of Carbohydrates Using the Hydroxyl Groups as Starting Points | ACS Omega